Gene name:
CDKN2A (CDKN2;MTS1) ;
Protein name:
Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3 ;
Alternative:
Multiple tumor suppressor 1 (MTS-1) ;Cyclin-dependent kinase 4 inhibitor A (CDK4I) ;p16-INK4a (p16-INK4;p16INK4A) ;
Organism:
Human (Homo sapiens).
General Annotation
Sub Unit:
Heterodimer with CDK4 or CDK6. Predominant p16 complexes contained CDK6. Interacts (isoforms 1,2 and 4) with CDK4 (both 'T-172'-phosphorylated and non-phosphorylated forms); the interaction inhibits cyclin D-CDK4 kinase activity. Interacts with ISCO2.
Function:
Acts as a negative regulator of the proliferation of normal cells by interacting strongly with CDK4 and CDK6. This inhibits their ability to interact with cyclins D and to phosphorylate the retinoblastoma protein.
Subcellular Location:
Cytoplasm
Nucleus
Protein Attributes:
50:
MEPAAGSSME | PSADWLATAA | ARGRVEEVRA | LLEAGALPNA | PNSYGRRPIQ |
100:
VMMMGSARVA | ELLLLHGAEP | NCADPATLTR | PVHDAAREGF | LDTLVVLHRA |
150:
GARLDVRDAW | GRLPVDLAEE | LGHRDVARYL | RAAAGGTRGS | NHARIDAAEG |
Vaild Sequence:
Related Databases
Uniprot:
ELISA Kit
CLIA Kit
Polyclonal Antibody
Monoclonal Antibody
Protein
FOR
Human
Rat
Mouse
ELISA Kit for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
ELISA Kit for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
ELISA Kit for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
CLIA Kit for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
CLIA Kit for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
CLIA Kit for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
Polyclonal Antibody for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
Polyclonal Antibody for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
Polyclonal Antibody for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
Monoclonal Antibody for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
Monoclonal Antibody for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
Monoclonal Antibody for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
Protein for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
Protein for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
Protein for Human Cyclin-dependent kinase inhibitor 2A, isoforms 1/2/3
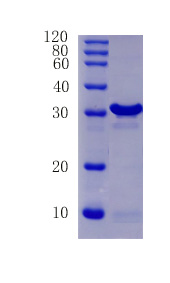
R&D Technical Data
12% SDS-PAGE Analysis
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
Precision
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
Recovery
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
Linearity
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
References
1.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [MRNA] (ISOFORM 1)
2.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [MRNA] (ISOFORM 3);TISSUE SPECIFICITY
3.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [GENOMIC DNA]
4.
"Human p16gamma, a novel transcriptional variant of p16(INK4A), coexpresses with p16(INK4A) in cancer cells and inhibits cell-cycle progression."
Lin Y.C.
,
Diccianni M.B.
,
Kim Y.
,
Lin H.H.
,
Lee C.H.
,
Lin R.J.
,
Joo S.H.
,
Li J.
,
Chuang T.J.
,
Yang A.S.
,
Kuo H.H.
,
Tsai M.D.
,
Yu A.L.
Oncogene26:7017-7027(2007)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [MRNA] (ISOFORM 5);ALTERNATIVE SPLICING
5.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [GENOMIC DNA]
6.
"DNA sequence and analysis of human chromosome 9."
Humphray S.J.
,
Oliver K.
,
Hunt A.R.
,
Plumb R.W.
,
Loveland J.E.
,
Howe K.L.
,
Andrews T.D.
,
Searle S.
,
Hunt S.E.
,
Scott C.E.
,
Jones M.C.
,
Ainscough R.
,
Almeida J.P.
,
Ambrose K.D.
,
Ashwell R.I.S.
,
Babbage A.K.
,
Babbage S.
,
Bagguley C.L.
,
Bailey J.
,
Banerjee R.
,
Barker D.J.
,
Barlow K.F.
,
Bates K.
,
Beasley H.
,
Beasley O.
,
Bird C.P.
,
Bray-Allen S.
,
Brown A.J.
,
Brown J.Y.
,
Burford D.
,
Burrill W.
,
Burton J.
,
Carder C.
,
Carter N.P.
,
Chapman J.C.
,
Chen Y.
,
Clarke G.
,
Clark S.Y.
,
Clee C.M.
,
Clegg S.
,
Collier R.E.
,
Corby N.
,
Crosier M.
,
Cummings A.T.
,
Davies J.
,
Dhami P.
,
Dunn M.
,
Dutta I.
,
Dyer L.W.
,
Earthrowl M.E.
,
Faulkner L.
,
Fleming C.J.
,
Frankish A.
,
Frankland J.A.
,
French L.
,
Fricker D.G.
,
Garner P.
,
Garnett J.
,
Ghori J.
,
Gilbert J.G.R.
,
Glison C.
,
Grafham D.V.
,
Gribble S.
,
Griffiths C.
,
Griffiths-Jones S.
,
Grocock R.
,
Guy J.
,
Hall R.E.
,
Hammond S.
,
Harley J.L.
,
Harrison E.S.I.
,
Hart E.A.
,
Heath P.D.
,
Henderson C.D.
,
Hopkins B.L.
,
Howard P.J.
,
Howden P.J.
,
Huckle E.
,
Johnson C.
,
Johnson D.
,
Joy A.A.
,
Kay M.
,
Keenan S.
,
Kershaw J.K.
,
Kimberley A.M.
,
King A.
,
Knights A.
,
Laird G.K.
,
Langford C.
,
Lawlor S.
,
Leongamornlert D.A.
,
Leversha M.
,
Lloyd C.
,
Lloyd D.M.
,
Lovell J.
,
Martin S.
,
Mashreghi-Mohammadi M.
,
Matthews L.
,
McLaren S.
,
McLay K.E.
,
McMurray A.
,
Milne S.
,
Nickerson T.
,
Nisbett J.
,
Nordsiek G.
,
Pearce A.V.
,
Peck A.I.
,
Porter K.M.
,
Pandian R.
,
Pelan S.
,
Phillimore B.
,
Povey S.
,
Ramsey Y.
,
Rand V.
,
Scharfe M.
,
Sehra H.K.
,
Shownkeen R.
,
Sims S.K.
,
Skuce C.D.
,
Smith M.
,
Steward C.A.
,
Swarbreck D.
,
Sycamore N.
,
Tester J.
,
Thorpe A.
,
Tracey A.
,
Tromans A.
,
Thomas D.W.
,
Wall M.
,
Wallis J.M.
,
West A.P.
,
Whitehead S.L.
,
Willey D.L.
,
Williams S.A.
,
Wilming L.
,
Wray P.W.
,
Young L.
,
Ashurst J.L.
,
Coulson A.
,
Blocker H.
,
Durbin R.M.
,
Sulston J.E.
,
Hubbard T.
,
Jackson M.J.
,
Bentley D.R.
,
Beck S.
,
Rogers J.
,
Dunham I.
more...
Nature429:369-374(2004)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [LARGE SCALE GENOMIC DNA]
7.
Mural R.J.
,
Istrail S.
,
Sutton G.G.
,
Florea L.
,
Halpern A.L.
,
Mobarry C.M.
,
Lippert R.
,
Walenz B.
,
Shatkay H.
,
Dew I.
,
Miller J.R.
,
Flanigan M.J.
,
Edwards N.J.
,
Bolanos R.
,
Fasulo D.
,
Halldorsson B.V.
,
Hannenhalli S.
,
Turner R.
,
Yooseph S.
,
Lu F.
,
Nusskern D.R.
,
Shue B.C.
,
Zheng X.H.
,
Zhong F.
,
Delcher A.L.
,
Huson D.H.
,
Kravitz S.A.
,
Mouchard L.
,
Reinert K.
,
Remington K.A.
,
Clark A.G.
,
Waterman M.S.
,
Eichler E.E.
,
Adams M.D.
,
Hunkapiller M.W.
,
Myers E.W.
,
Venter J.C.
more...
Submitted (2005-09) to the EMBL/GenBank/DDBJ databases
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [LARGE SCALE GENOMIC DNA]
8.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [GENOMIC DNA] OF 1-20
9.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [GENOMIC DNA] OF 51-156
10.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [GENOMIC DNA] OF 51-152
11.
"Mutations and altered expression of p16INK4 in human cancer."
Okamoto A.
,
Demetrick D.J.
,
Spillare E.A.
,
Hagiwara K.
,
Hussain S.P.
,
Bennett W.P.
,
Forrester K.
,
Gerwin B.
,
Serrano M.
,
Beach D.H.
,
Harris C.C.
Proc. Natl. Acad. Sci. U.S.A.91:11045-11049(1994)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for FUNCTION
12.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for PHOSPHORYLATION AT SER-7; SER-8; SER-140 AND SER-152
13.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for INTERACTION WITH CDK4;FUNCTION
14.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for INTERACTION WITH ISOC2;SUBCELLULAR LOCATION
15.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for ACETYLATION [LARGE SCALE ANALYSIS] AT MET-1;IDENTIFICATION BY MASS SPECTROMETRY [LARGE SCALE ANALYSIS]
16.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for IDENTIFICATION BY MASS SPECTROMETRY [LARGE SCALE ANALYSIS]
17.
"N-terminal acetylome analyses and functional insights of the N-terminal acetyltransferase NatB."
Van Damme P.
,
Lasa M.
,
Polevoda B.
,
Gazquez C.
,
Elosegui-Artola A.
,
Kim D.S.
,
De Juan-Pardo E.
,
Demeyer K.
,
Hole K.
,
Larrea E.
,
Timmerman E.
,
Prieto J.
,
Arnesen T.
,
Sherman F.
,
Gevaert K.
,
Aldabe R.
Proc. Natl. Acad. Sci. U.S.A.109:12449-12454(2012)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for ACETYLATION [LARGE SCALE ANALYSIS] AT MET-1;IDENTIFICATION BY MASS SPECTROMETRY [LARGE SCALE ANALYSIS]
18.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for X-RAY CRYSTALLOGRAPHY (2.8 ANGSTROMS) OF COMPLEX WITH CDK6
19.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for STRUCTURE BY NMR
20.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for STRUCTURE BY NMR
21.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for REVIEW ON MELANOMA VARIANTS
22.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for REVIEW ON VARIANTS
23.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS NON-SMALL CELL LUNG CARCINOMA TYR-66; HIS-84; ALA-93; ALA-95; GLN-99; LEU-114; ALA-120; LYS-120; PRO-132; VAL-134; TYR-142 AND VAL-150
24.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS CMM2 PRO-87; TRP-101 AND ASP-126;VARIANTS THR-49; SER-71 AND THR-148
25.
"Prevalence of germ-line mutations in p16, p19ARF, and CDK4 in familial melanoma: analysis of a clinic-based population."
Fitzgerald M.G.
,
Harkin D.P.
,
Silva-Arrieta S.
,
Macdonald D.J.
,
Lucchina L.C.
,
Unsal H.
,
O'Neill E.
,
Koh J.
,
Finkelstein D.M.
,
Isselbacher K.J.
,
Sober A.J.
,
Haber D.A.
Proc. Natl. Acad. Sci. U.S.A.93:8541-8545(1996)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS CMM2 ILE-53 AND CYS-107;VARIANTS VAL-68; THR-85 AND THR-148
26.
"Germline mutations of the CDKN2 gene in UK melanoma families."
Harland M.
,
Meloni R.
,
Gruis N.
,
Pinney E.
,
Brookes S.
,
Spurr N.K.
,
Frischauf A.-M.
,
Bataille V.
,
Peters G.
,
Cuzick J.
,
Selby P.
,
Bishop D.T.
,
Bishop J.N.
Hum. Mol. Genet.6:2061-2067(1997)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS CMM2 PRO-24; ILE-53 AND THR-118;VARIANT THR-148
27.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS CMM2
28.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for ERRATUM
29.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS CMM2 LEU-48; ASP-89 AND MET-117;VARIANT PANCREAS CARCINOMA VAL-57;VARIANT THR-148
30.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANT LFS GLU-102
31.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANT CMM2 ASP-126
32.
"A germline deletion of p14(ARF) but not CDKN2A in a melanoma-neural system tumour syndrome family."
Randerson-Moor J.A.
,
Harland M.
,
Williams S.
,
Cuthbert-Heavens D.
,
Sheridan E.
,
Aveyard J.
,
Sibley K.
,
Whitaker L.
,
Knowles M.
,
Bishop J.N.
,
Bishop D.T.
Hum. Mol. Genet.10:55-62(2001)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for INVOLVEMENT IN MASTS
33.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS SQUAMOUS CELL CARCINOMA SER-127 AND CYS-144
34.
"Mutations in the p16INK4/MTS1/CDKN2, p15INK4B/MTS2, and p18 genes in primary and metastatic lung cancer."
Okamoto A.
,
Hussain S.P.
,
Hagiwara K.
,
Spillare E.A.
,
Rusin M.R.
,
Demetrick D.J.
,
Serrano M.
,
Hannon G.J.
,
Shiseki M.
,
Zariwala M.
,
Xiong Y.
,
Beach D.H.
,
Yokota J.
,
Harris C.C.
Cancer Res.55:1448-1451(1995)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS
35.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS CMM2 PRO-32; ALA-35; GLU-35; ARG-50 AND ILE-53;VARIANT THR-148
36.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for CHARACTERIZATION OF VARIANTS THR-49; SER-71; LEU-81; PRO-87; TRP-101; ASP-126 AND THR-148
37.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANT CMM2 ARG-112 INS;VARIANT THR-148
38.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANT CMM2 ARG-122
39.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS CMM2 GLY-59; TYR-84; TRP-87 AND TRP-101
40.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for POSSIBLE INVOLVEMENT IN SUSCEPTIBILITY TO UVEAL MELANOMA
41.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANT CMM2 GLN-94
42.
"Functional, structural, and genetic evaluation of 20 CDKN2A germ line mutations identified in melanoma-prone families or patients."
Kannengiesser C.
,
Brookes S.
,
del Arroyo A.G.
,
Pham D.
,
Bombled J.
,
Barrois M.
,
Mauffret O.
,
Avril M.F.
,
Chompret A.
,
Lenoir G.M.
,
Sarasin A.
,
Peters G.
,
Bressac-de Paillerets B.
Hum. Mutat.30:564-574(2009)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS CMM2 THR-ALA-19 INS; VAL-35; ARG-67; 67-GLY--ASN-71 DEL; TYR-74; PRO-77; PRO-80; THR-81 AND ARG-97;VARIANTS GLN-24; GLY-69 AND SER-114;CHARACTERIZATION OF VARIANTS