Protein name:
Endoglin ;
Alternative:
CD:
CD105 ;
Organism:
Human (Homo sapiens).
General Annotation
Sub Unit:
Homodimer that forms an heteromeric complex with the signaling receptors for transforming growth factor-beta: TGFBR1 and/or TGFBR2. It is able to bind TGF-beta 1, and 3 efficiently and TGF-beta 2 less efficiently. Interacts with TCTEX1D4. Interacts with ARRB2.
Function:
Major glycoprotein of vascular endothelium. May play a critical role in the binding of endothelial cells to integrins and/or other RGD receptors.
Subcellular Location:
Membrane
Single-pass type I membrane protein
Protein Attributes:
50:
MDRGTLPLAV | ALLLASCSLS | PTSLAETVHC | DLQPVGPERG | EVTYTTSQVS |
100:
KGCVAQAPNA | ILEVHVLFLE | FPTGPSQLEL | TLQASKQNGT | WPREVLLVLS |
150:
VNSSVFLHLQ | ALGIPLHLAY | NSSLVTFQEP | PGVNTTELPS | FPKTQILEWA |
200:
AERGPITSAA | ELNDPQSILL | RLGQAQGSLS | FCMLEASQDM | GRTLEWRPRT |
250:
PALVRGCHLE | GVAGHKEAHI | LRVLPGHSAG | PRTVTVKVEL | SCAPGDLDAV |
300:
LILQGPPYVS | WLIDANHNMQ | IWTTGEYSFK | IFPEKNIRGF | KLPDTPQGLL |
350:
GEARMLNASI | VASFVELPLA | SIVSLHASSC | GGRLQTSPAP | IQTTPPKDTC |
400:
SPELLMSLIQ | TKCADDAMTL | VLKKELVAHL | KCTITGLTFW | DPSCEAEDRG |
450:
DKFVLRSAYS | SCGMQVSASM | ISNEAVVNIL | SSSSPQRKKV | HCLNMDSLSF |
500:
QLGLYLSPHF | LQASNTIEPG | QQSFVQVRVS | PSVSEFLLQL | DSCHLDLGPE |
550:
GGTVELIQGR | AAKGNCVSLL | SPSPEGDPRF | SFLLHFYTVP | IPKTGTLSCT |
600:
VALRPKTGSQ | DQEVHRTVFM | RLNIISPDLS | GCTSKGLVLP | AVLGITFGAF |
650:
LIGALLTAAL | WYIYSHTRSP | SKREPVVAVA | APASSESSST | NHSIGSTQST |
Vaild Sequence:
Related Databases
Uniprot:
ELISA Kit
CLIA Kit
Polyclonal Antibody
Monoclonal Antibody
Protein
FOR
Mouse
Human
Pig
ELISA Kit for Human Endoglin
ELISA Kit for Human Endoglin
ELISA Kit for Human Endoglin
CLIA Kit for Human Endoglin
CLIA Kit for Human Endoglin
CLIA Kit for Human Endoglin
Polyclonal Antibody for Human Endoglin
Polyclonal Antibody for Human Endoglin
Polyclonal Antibody for Human Endoglin
Monoclonal Antibody for Human Endoglin
Monoclonal Antibody for Human Endoglin
Monoclonal Antibody for Human Endoglin
Protein for Human Endoglin
Protein for Human Endoglin
Protein for Human Endoglin
R&D Technical Data
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
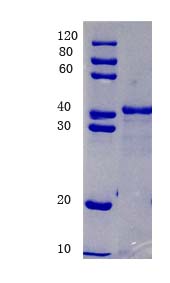
12% SDS-PAGE analysis
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
Precision
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
Recovery
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
Linearity
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
For more information, please refer to the manual,Or contact our technical support: tech@eiaab.com.
References
1.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [MRNA] (ISOFORM SHORT)
2.
"DNA sequence and analysis of human chromosome 9."
Humphray S.J.
,
Oliver K.
,
Hunt A.R.
,
Plumb R.W.
,
Loveland J.E.
,
Howe K.L.
,
Andrews T.D.
,
Searle S.
,
Hunt S.E.
,
Scott C.E.
,
Jones M.C.
,
Ainscough R.
,
Almeida J.P.
,
Ambrose K.D.
,
Ashwell R.I.S.
,
Babbage A.K.
,
Babbage S.
,
Bagguley C.L.
,
Bailey J.
,
Banerjee R.
,
Barker D.J.
,
Barlow K.F.
,
Bates K.
,
Beasley H.
,
Beasley O.
,
Bird C.P.
,
Bray-Allen S.
,
Brown A.J.
,
Brown J.Y.
,
Burford D.
,
Burrill W.
,
Burton J.
,
Carder C.
,
Carter N.P.
,
Chapman J.C.
,
Chen Y.
,
Clarke G.
,
Clark S.Y.
,
Clee C.M.
,
Clegg S.
,
Collier R.E.
,
Corby N.
,
Crosier M.
,
Cummings A.T.
,
Davies J.
,
Dhami P.
,
Dunn M.
,
Dutta I.
,
Dyer L.W.
,
Earthrowl M.E.
,
Faulkner L.
,
Fleming C.J.
,
Frankish A.
,
Frankland J.A.
,
French L.
,
Fricker D.G.
,
Garner P.
,
Garnett J.
,
Ghori J.
,
Gilbert J.G.R.
,
Glison C.
,
Grafham D.V.
,
Gribble S.
,
Griffiths C.
,
Griffiths-Jones S.
,
Grocock R.
,
Guy J.
,
Hall R.E.
,
Hammond S.
,
Harley J.L.
,
Harrison E.S.I.
,
Hart E.A.
,
Heath P.D.
,
Henderson C.D.
,
Hopkins B.L.
,
Howard P.J.
,
Howden P.J.
,
Huckle E.
,
Johnson C.
,
Johnson D.
,
Joy A.A.
,
Kay M.
,
Keenan S.
,
Kershaw J.K.
,
Kimberley A.M.
,
King A.
,
Knights A.
,
Laird G.K.
,
Langford C.
,
Lawlor S.
,
Leongamornlert D.A.
,
Leversha M.
,
Lloyd C.
,
Lloyd D.M.
,
Lovell J.
,
Martin S.
,
Mashreghi-Mohammadi M.
,
Matthews L.
,
McLaren S.
,
McLay K.E.
,
McMurray A.
,
Milne S.
,
Nickerson T.
,
Nisbett J.
,
Nordsiek G.
,
Pearce A.V.
,
Peck A.I.
,
Porter K.M.
,
Pandian R.
,
Pelan S.
,
Phillimore B.
,
Povey S.
,
Ramsey Y.
,
Rand V.
,
Scharfe M.
,
Sehra H.K.
,
Shownkeen R.
,
Sims S.K.
,
Skuce C.D.
,
Smith M.
,
Steward C.A.
,
Swarbreck D.
,
Sycamore N.
,
Tester J.
,
Thorpe A.
,
Tracey A.
,
Tromans A.
,
Thomas D.W.
,
Wall M.
,
Wallis J.M.
,
West A.P.
,
Whitehead S.L.
,
Willey D.L.
,
Williams S.A.
,
Wilming L.
,
Wray P.W.
,
Young L.
,
Ashurst J.L.
,
Coulson A.
,
Blocker H.
,
Durbin R.M.
,
Sulston J.E.
,
Hubbard T.
,
Jackson M.J.
,
Bentley D.R.
,
Beck S.
,
Rogers J.
,
Dunham I.
more...
Nature429:369-374(2004)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [LARGE SCALE GENOMIC DNA]
3.
Mural R.J.
,
Istrail S.
,
Sutton G.G.
,
Florea L.
,
Halpern A.L.
,
Mobarry C.M.
,
Lippert R.
,
Walenz B.
,
Shatkay H.
,
Dew I.
,
Miller J.R.
,
Flanigan M.J.
,
Edwards N.J.
,
Bolanos R.
,
Fasulo D.
,
Halldorsson B.V.
,
Hannenhalli S.
,
Turner R.
,
Yooseph S.
,
Lu F.
,
Nusskern D.R.
,
Shue B.C.
,
Zheng X.H.
,
Zhong F.
,
Delcher A.L.
,
Huson D.H.
,
Kravitz S.A.
,
Mouchard L.
,
Reinert K.
,
Remington K.A.
,
Clark A.G.
,
Waterman M.S.
,
Eichler E.E.
,
Adams M.D.
,
Hunkapiller M.W.
,
Myers E.W.
,
Venter J.C.
more...
Submitted (2005-07) to the EMBL/GenBank/DDBJ databases
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [LARGE SCALE GENOMIC DNA]
4.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [GENOMIC DNA / MRNA] OF 14-658;PROTEIN SEQUENCE OF 26-36 (ISOFORM LONG)
tissue :
Umbilical vein .
5.
"Endoglin, a TGF-beta binding protein of endothelial cells, is the gene for hereditary haemorrhagic telangiectasia type 1."
McAllister K.A.
,
Grogg K.M.
,
Johnson D.W.
,
Gallione C.J.
,
Baldwin M.A.
,
Jackson C.E.
,
Helmbold E.A.
,
Markel D.S.
,
McKinnon W.C.
,
Murrell J.
,
McCormick M.K.
,
Pericak-Vance M.A.
,
Heutink P.
,
Oostra B.A.
,
Haitjema T.
,
Westerman C.J.
,
Porteous M.E.
,
Guttmacher A.E.
,
Letarte M.
,
Marchuk D.A.
more...
Nat. Genet.8:345-351(1994)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for NUCLEOTIDE SEQUENCE [GENOMIC DNA] OF 122-378
6.
"Identification of Tctex2beta, a novel dynein light chain family member that interacts with different transforming growth factor-beta receptors."
Meng Q.-J.
,
Lux A.
,
Holloschi A.
,
Li J.
,
Hughes J.M.X.
,
Foerg T.
,
McCarthy J.E.G.
,
Heagerty A.M.
,
Kioschis P.
,
Hafner M.
,
Garland J.M.
J. Biol. Chem.281:37069-37080(2006)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for INTERACTION WITH TCTEX1D4
7.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for INTERACTION WITH ARRB2
8.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for GLYCOSYLATION [LARGE SCALE ANALYSIS] AT ASN-134
tissue :
Liver .
9.
"Soluble endoglin specifically binds bone morphogenetic proteins 9 and 10 via its orphan domain, inhibits blood vessel formation, and suppresses tumor growth."
Castonguay R.
,
Werner E.D.
,
Matthews R.G.
,
Presman E.
,
Mulivor A.W.
,
Solban N.
,
Sako D.
,
Pearsall R.S.
,
Underwood K.W.
,
Seehra J.
,
Kumar R.
,
Grinberg A.V.
J. Biol. Chem.286:30034-30046(2011)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for FUNCTION;INTERACTION WITH GDF2
10.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for INTERACTION WITH GDF2
11.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for FUNCTION
12.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANT HHT1 192-ARG--PRO-198 DEL;VARIANT MET-5
13.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANT HHT1 ASP-160
14.
"Mutation and expression analysis of the endoglin gene in hereditary hemorrhagic telangiectasia reveals null alleles."
Gallione C.J.
,
Klaus D.J.
,
Yeh E.Y.
,
Stenzel T.T.
,
Xue Y.
,
Anthony K.B.
,
McAllister K.A.
,
Baldwin M.A.
,
Berg J.N.
,
Lux A.
,
Smith J.D.
,
Vary C.P.H.
,
Craigen W.J.
,
Westermann C.J.J.
,
Warner M.L.
,
Miller Y.E.
,
Jackson C.E.
,
Guttmacher A.E.
,
Marchuk D.A.
more...
Hum. Mutat.11:286-294(1998)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS HHT1 VAL-52; ARG-53; CYS-149 AND PRO-306
15.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS HHT1 VAL-52; ARG-53; CYS-149 AND PRO-221
16.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANT HHT1 VAL-413
17.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANT HHT1 ARG-53
18.
"Molecular screening of ALK1/ACVRL1 and ENG genes in hereditary hemorrhagic telangiectasia in France."
French Rendu-Osler network
,
Lesca G.
,
Plauchu H.
,
Coulet F.
,
Lefebvre S.
,
Plessis G.
,
Odent S.
,
Riviere S.
,
Leheup B.
,
Goizet C.
,
Carette M.-F.
,
Cordier J.-F.
,
Pinson S.
,
Soubrier F.
,
Calender A.
,
Giraud S.
Hum. Mutat.23:289-299(2004)
[
PubMed ]
[
Europe PMC ]
[
Abstract ]
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS HHT1 PRO-8; PHE-49; ARG-107; CYS-207 DEL; THR-263; ARG-232-233-THR DEL; ILE-263 DEL; SER-412 AND MET-504
19.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS HHT1 PRO-221; ILE-263 DEL AND LEU-615
20.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS HHT1 ASP-11; ASP-105; GLU-175; THR-220; ASP-308; SER-363; TRP-437; SER-490; HIS-529; PRO-547 AND ASP-604
21.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS HHT1 193-THR-LEU-194 DELINS VAL-LEU-GLN AND ASP-545
22.
[15/1/25 17:38] Upload to ab completed in less than a minute: 1 file transferred (13.4 Kb/s)
Cited for VARIANTS PRO-150; PRO-205; MET-236; MET-315; GLU-374; ARG-414; SER-545; TYR-549 AND ALA-561;VARIANTS HHT1 GLN-221; GLU-238; SER-263; ARG-269; TYR-394; PRO-529 AND ARG-603